Statistical Analysis in R
Last updated on 2026-04-22 | Edit this page
Estimated time: 120 minutes
Overview
Questions
- How do I run t-tests, ANOVA, correlations, and regression in R?
- How does R output compare to SPSS output tables?
- How do I extract and report results?
Objectives
- Run independent and paired samples t-tests
- Conduct one-way ANOVA with post-hoc tests
- Calculate correlations and run simple linear regression
- Interpret R output by mapping it to familiar SPSS output tables

From SPSS dialogs to R functions
In SPSS, every statistical test lives behind a menu: Analyze > Compare Means, Analyze > Correlate, and so on. In R, each test is a single function call. The table below maps the SPSS dialogs you already know to their R equivalents:
| Analysis | SPSS menu path | R function |
|---|---|---|
| Independent-samples t-test | Analyze > Compare Means > Independent-Samples T Test | t.test(y ~ group, data = df) |
| Paired-samples t-test | Analyze > Compare Means > Paired-Samples T Test | t.test(x, y, paired = TRUE) |
| One-way ANOVA | Analyze > Compare Means > One-Way ANOVA |
aov(y ~ group, data = df) + summary()
|
| Post-hoc tests | (checkbox in ANOVA dialog) | TukeyHSD() |
| Bivariate correlation | Analyze > Correlate > Bivariate | cor.test(df$x, df$y) |
| Linear regression | Analyze > Regression > Linear |
lm(y ~ x1 + x2, data = df) +
summary()
|
Let’s load our data and packages:
R
library(tidyverse)
library(broom)
visitors <- read_csv("data/aruba_visitors.csv")
Checking normality
Before running parametric tests like the t-test or ANOVA, you need to assess whether your data are approximately normally distributed. In SPSS, you would go to Analyze > Descriptive Statistics > Explore, move your variable to the “Dependent List”, click “Plots”, and check the “Normality plots with tests” box. R gives you the same tools – and more control over how they look.
Visual inspection: histogram and Q-Q plot
You already know ggplot2 from Episode 04, so let’s use
it. A histogram shows the shape of the distribution, and a Q-Q
(quantile-quantile) plot compares your data to a theoretical normal
distribution. If the points fall roughly along the diagonal line, the
data are approximately normal.
R
# Iteration: 1
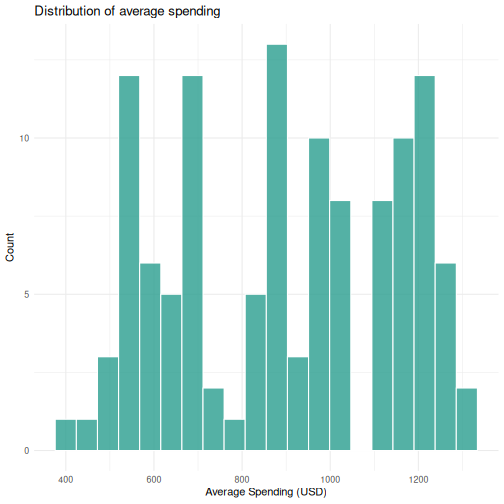
# Histogram of average spending
ggplot(visitors, aes(x = avg_spending_usd)) +
geom_histogram(bins = 20, fill = "#2a9d8f", color = "white", alpha = 0.8) +
labs(title = "Distribution of average spending",
x = "Average Spending (USD)",
y = "Count") +
theme_minimal()
R
# Iteration: 1
# Q-Q plot of average spending
ggplot(visitors, aes(sample = avg_spending_usd)) +
stat_qq() +
stat_qq_line(color = "#e76f51", linewidth = 0.8) +
labs(title = "Q-Q plot of average spending",
x = "Theoretical quantiles",
y = "Sample quantiles") +
theme_minimal()
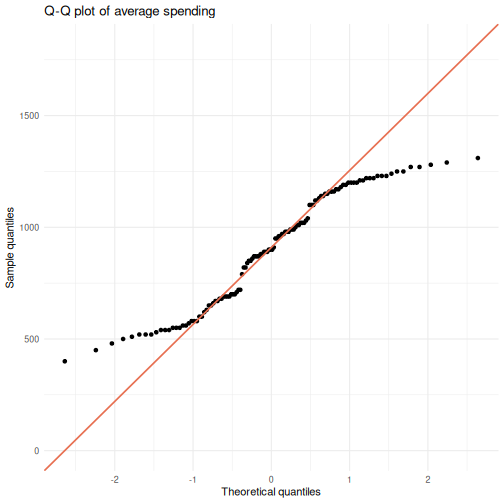
Points that hug the diagonal line suggest normality. Systematic deviations – S-curves, banana shapes, or points flying off at the tails – indicate non-normality.
Shapiro-Wilk test
The Shapiro-Wilk test is the formal statistical test for normality. The null hypothesis is that the data are normally distributed, so a significant p-value (< 0.05) means you reject normality.
R
# Iteration: 1
shapiro.test(visitors$avg_spending_usd)
OUTPUT
Shapiro-Wilk normality test
data: visitors$avg_spending_usd
W = 0.93876, p-value = 3.534e-05In SPSS, this same test appears in the “Tests of Normality” table when you check the normality plots option in Explore.
When does normality actually matter?
Parametric tests like the t-test and ANOVA assume normality, but they are surprisingly robust to violations of this assumption – especially with larger samples. As a rough guide:
- With n > 30 per group, the Central Limit Theorem means the sampling distribution of the mean is approximately normal regardless of the data distribution. The t-test and ANOVA will perform well.
- With small samples (n < 15 per group), normality matters more. Consider non-parametric alternatives (Wilcoxon test, Kruskal-Wallis) if the data are clearly non-normal.
- Always look at the plots first. The Shapiro-Wilk test is sensitive to trivial deviations in large samples, so a significant p-value does not necessarily mean parametric tests are inappropriate.
The combination of visual inspection and the formal test gives you a complete picture – far more useful than relying on either one alone.
Challenge: Check normality for satisfaction scores
- Create a histogram of
satisfaction_scoreusingggplot2. - Create a Q-Q plot of
satisfaction_score. - Run the Shapiro-Wilk test on
satisfaction_score. - Based on all three, would you feel comfortable running a parametric test on this variable? Why or why not?
R
# Iteration: 1
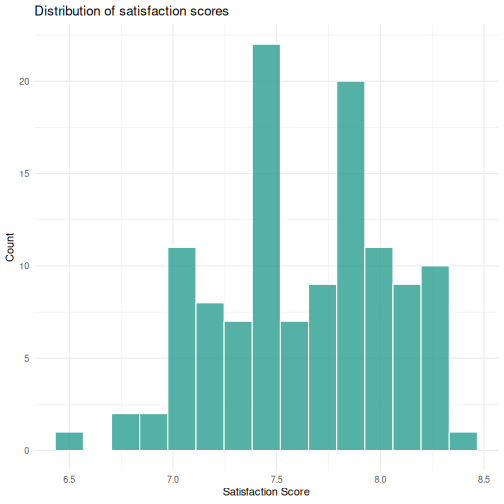
# 1: Histogram
ggplot(visitors, aes(x = satisfaction_score)) +
geom_histogram(bins = 15, fill = "#2a9d8f", color = "white", alpha = 0.8) +
labs(title = "Distribution of satisfaction scores",
x = "Satisfaction Score",
y = "Count") +
theme_minimal()
R
# 2: Q-Q plot
ggplot(visitors, aes(sample = satisfaction_score)) +
stat_qq() +
stat_qq_line(color = "#e76f51", linewidth = 0.8) +
labs(title = "Q-Q plot of satisfaction scores",
x = "Theoretical quantiles",
y = "Sample quantiles") +
theme_minimal()
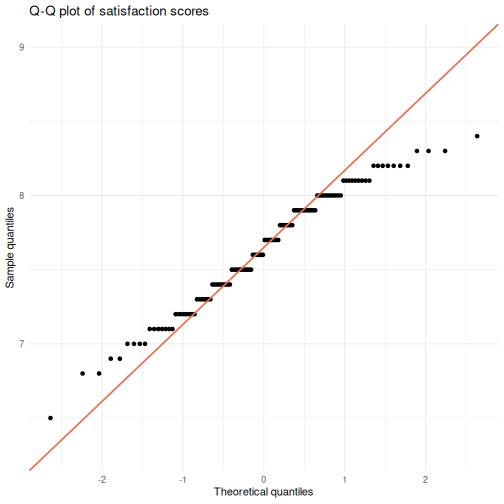
R
# 3: Shapiro-Wilk test
shapiro.test(visitors$satisfaction_score)
OUTPUT
Shapiro-Wilk normality test
data: visitors$satisfaction_score
W = 0.97421, p-value = 0.02086R
# 4: Interpretation depends on the output. With n = 120, the t-test is
# robust to moderate non-normality. Check whether the histogram looks
# roughly symmetric and the Q-Q points stay close to the line.
T-tests
Independent-samples t-test
In SPSS, you would go to Analyze > Compare Means > Independent-Samples T Test, move your test variable to the “Test Variable(s)” box, move your grouping variable to the “Grouping Variable” box, and define the two groups.
In R, it is one line. Let’s test whether the average spending differs between US and Netherlands visitors:
R
# Filter to just the two groups we want to compare
us_nl <- visitors |>
filter(origin %in% c("United States", "Netherlands"))
# Run the independent-samples t-test
t_result <- t.test(avg_spending_usd ~ origin, data = us_nl)
t_result
OUTPUT
Welch Two Sample t-test
data: avg_spending_usd by origin
t = -16.195, df = 37.99, p-value < 2.2e-16
alternative hypothesis: true difference in means between group Netherlands and group United States is not equal to 0
95 percent confidence interval:
-280.1256 -217.8744
sample estimates:
mean in group Netherlands mean in group United States
978.5 1227.5 Reading the t-test output — SPSS comparison
The R output gives you the same information as the SPSS “Independent Samples Test” table, just arranged differently:
| SPSS output column | R output line |
|---|---|
| t | t = ... |
| df | df = ... |
| Sig. (2-tailed) | p-value = ... |
| Mean Difference | difference in means shown in estimates |
| 95% CI of the Difference | 95 percent confidence interval: |
The key difference: SPSS shows Levene’s test for equality of
variances automatically. R’s t.test() uses the Welch
correction by default (which does not assume equal
variances). This is actually the better default — many statisticians
recommend always using the Welch t-test.
If you need the equal-variances version (the “Equal variances
assumed” row in SPSS), add var.equal = TRUE:
R
t.test(avg_spending_usd ~ origin, data = us_nl, var.equal = TRUE)
Paired-samples t-test
A paired t-test compares two measurements on the same cases. Let’s compare Q1 vs. Q3 hotel occupancy rates (the same hotels measured in different quarters):
R
# Get Q1 and Q3 data for each year
q1_data <- visitors |>
filter(quarter == "Q1") |>
arrange(year, origin) |>
pull(hotel_occupancy_pct)
q3_data <- visitors |>
filter(quarter == "Q3") |>
arrange(year, origin) |>
pull(hotel_occupancy_pct)
# Paired t-test
t.test(q1_data, q3_data, paired = TRUE)
OUTPUT
Paired t-test
data: q1_data and q3_data
t = 3.6602, df = 29, p-value = 0.000998
alternative hypothesis: true mean difference is not equal to 0
95 percent confidence interval:
4.941648 17.458352
sample estimates:
mean difference
11.2 In SPSS this would be Analyze > Compare Means > Paired-Samples T Test, where you select the two variables as a pair.
ANOVA
One-way ANOVA
In SPSS: Analyze > Compare Means > One-Way ANOVA. Move the dependent variable to the “Dependent List” and the factor to “Factor”.
Let’s test whether satisfaction scores differ across origin countries:
R
# Fit the ANOVA model
anova_model <- aov(satisfaction_score ~ origin, data = visitors)
# View the ANOVA table
summary(anova_model)
OUTPUT
Df Sum Sq Mean Sq F value Pr(>F)
origin 5 11.610 2.3220 35.28 <2e-16 ***
Residuals 114 7.502 0.0658
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Reading the ANOVA table — SPSS comparison
| SPSS output column | R output column |
|---|---|
| Sum of Squares | Sum Sq |
| df | Df |
| Mean Square | Mean Sq |
| F | F value |
| Sig. | Pr(>F) |
The layout is almost identical — R just uses slightly different column names.
Post-hoc tests with TukeyHSD
If the ANOVA is significant, you want to know which groups differ. In SPSS, you check the “Post Hoc” button in the ANOVA dialog and select Tukey. In R:
R
TukeyHSD(anova_model)
OUTPUT
Tukey multiple comparisons of means
95% family-wise confidence level
Fit: aov(formula = satisfaction_score ~ origin, data = visitors)
$origin
diff lwr upr p adj
Colombia-Canada -0.495 -0.7301532 -0.25984681 0.0000002
Netherlands-Canada -0.020 -0.2551532 0.21515319 0.9998728
Other-Canada -0.420 -0.6551532 -0.18484681 0.0000143
United States-Canada 0.100 -0.1351532 0.33515319 0.8199046
Venezuela-Canada -0.755 -0.9901532 -0.51984681 0.0000000
Netherlands-Colombia 0.475 0.2398468 0.71015319 0.0000007
Other-Colombia 0.075 -0.1601532 0.31015319 0.9394347
United States-Colombia 0.595 0.3598468 0.83015319 0.0000000
Venezuela-Colombia -0.260 -0.4951532 -0.02484681 0.0211495
Other-Netherlands -0.400 -0.6351532 -0.16484681 0.0000408
United States-Netherlands 0.120 -0.1151532 0.35515319 0.6781085
Venezuela-Netherlands -0.735 -0.9701532 -0.49984681 0.0000000
United States-Other 0.520 0.2848468 0.75515319 0.0000001
Venezuela-Other -0.335 -0.5701532 -0.09984681 0.0009635
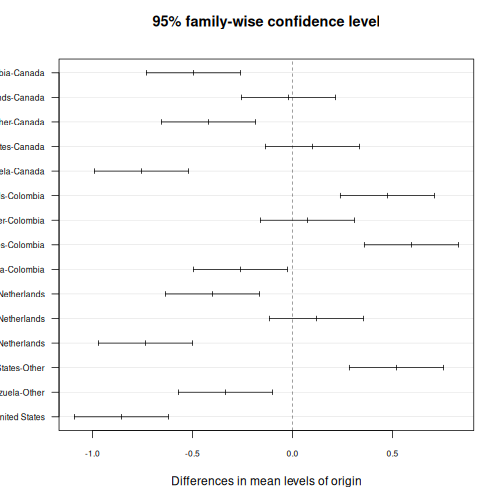
Venezuela-United States -0.855 -1.0901532 -0.61984681 0.0000000This gives you pairwise comparisons with adjusted p-values, the difference in means, and 95% confidence intervals — the same information as the SPSS “Multiple Comparisons” table.
You can also visualize the post-hoc results:
R
plot(TukeyHSD(anova_model), las = 1, cex.axis = 0.7)
Correlation
In SPSS: Analyze > Correlate > Bivariate. Move variables to the “Variables” box and select Pearson, Spearman, or both.
Let’s test the correlation between average spending and satisfaction:
R
cor.test(visitors$avg_spending_usd, visitors$satisfaction_score)
OUTPUT
Pearson's product-moment correlation
data: visitors$avg_spending_usd and visitors$satisfaction_score
t = 18.69, df = 118, p-value < 2.2e-16
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
0.8110127 0.9037614
sample estimates:
cor
0.8645736 Reading correlation output — SPSS comparison
| SPSS output | R output |
|---|---|
| Pearson Correlation |
cor (the estimate at the end) |
| Sig. (2-tailed) | p-value |
| N | shown in the data, not in output |
| 95% CI | 95 percent confidence interval |
One advantage of R: cor.test() gives you a confidence
interval for the correlation by default. SPSS does not show this unless
you use syntax.
For a correlation matrix of multiple variables (like the SPSS
correlation table), use cor():
R
visitors |>
select(visitors_stayover, avg_stay_nights, avg_spending_usd,
hotel_occupancy_pct, satisfaction_score) |>
cor(use = "complete.obs") |>
round(3)
OUTPUT
visitors_stayover avg_stay_nights avg_spending_usd
visitors_stayover 1.000 0.276 0.622
avg_stay_nights 0.276 1.000 0.597
avg_spending_usd 0.622 0.597 1.000
hotel_occupancy_pct 0.231 0.166 0.206
satisfaction_score 0.589 0.674 0.865
hotel_occupancy_pct satisfaction_score
visitors_stayover 0.231 0.589
avg_stay_nights 0.166 0.674
avg_spending_usd 0.206 0.865
hotel_occupancy_pct 1.000 0.575
satisfaction_score 0.575 1.000Chi-square test of independence
All of the tests above work with numeric variables. When both variables are categorical, you need a different tool: the chi-square test of independence. In SPSS: Analyze > Descriptive Statistics > Crosstabs, move one variable to “Row(s)” and another to “Column(s)”, then click “Statistics” and check the “Chi-square” box.
Building a contingency table
First, create a contingency table using table(). Let’s
test whether the distribution of visitor origin countries
differs across quarter:
R
# Iteration: 1
# Create the contingency table
origin_quarter <- table(visitors$origin, visitors$quarter)
origin_quarter
OUTPUT
Q1 Q2 Q3 Q4
Canada 5 5 5 5
Colombia 5 5 5 5
Netherlands 5 5 5 5
Other 5 5 5 5
United States 5 5 5 5
Venezuela 5 5 5 5This is the same table you would see in the “Crosstabulation” output in SPSS. Each cell shows the observed count for that combination of origin and quarter.
Running the chi-square test
R
# Iteration: 1
chi_result <- chisq.test(origin_quarter)
chi_result
OUTPUT
Pearson's Chi-squared test
data: origin_quarter
X-squared = 0, df = 15, p-value = 1The output gives you three pieces of information, matching the SPSS “Chi-Square Tests” table:
| SPSS output column | R output |
|---|---|
| Pearson Chi-Square value | X-squared |
| df | df |
| Asymp. Sig. (2-sided) | p-value |
A significant p-value means the distribution of one variable depends on the other – the row and column variables are not independent.
Expected vs observed frequencies
The expected frequencies tell you what the cell counts would look like if the two variables were completely independent. Comparing expected to observed is how you identify which cells are driving the result.
R
# Iteration: 1
# Expected frequencies (what you'd see under independence)
chi_result$expected |> round(1)
OUTPUT
Q1 Q2 Q3 Q4
Canada 5 5 5 5
Colombia 5 5 5 5
Netherlands 5 5 5 5
Other 5 5 5 5
United States 5 5 5 5
Venezuela 5 5 5 5R
# Iteration: 1
# Standardized residuals -- large absolute values (> 2) flag notable cells
chi_result$residuals |> round(2)
OUTPUT
Q1 Q2 Q3 Q4
Canada 0 0 0 0
Colombia 0 0 0 0
Netherlands 0 0 0 0
Other 0 0 0 0
United States 0 0 0 0
Venezuela 0 0 0 0In SPSS, you would get these by checking “Expected” and “Standardized” under the “Cells” button in the Crosstabs dialog. Residuals greater than 2 or less than -2 point to cells where observed counts deviate substantially from what independence predicts.
Challenge: Chi-square analysis
Create a new categorical variable called spending_level
that classifies rows as “High” (above the median of
avg_spending_usd) or “Low” (at or below the median). Then
test whether spending_level is independent of
origin.
- Create
spending_levelusingmutate()andif_else(). - Build a contingency table of
originbyspending_level. - Run
chisq.test()and interpret the result. - Examine the standardized residuals to see which origin countries spend more (or less) than expected.
R
# Iteration: 1
# 1: Create spending_level
visitors_spending <- visitors |>
mutate(spending_level = if_else(
avg_spending_usd > median(avg_spending_usd), "High", "Low"
))
# 2: Contingency table
spending_table <- table(visitors_spending$origin,
visitors_spending$spending_level)
spending_table
OUTPUT
High Low
Canada 20 0
Colombia 0 20
Netherlands 18 2
Other 1 19
United States 20 0
Venezuela 0 20R
# 3: Chi-square test
chi_spending <- chisq.test(spending_table)
chi_spending
OUTPUT
Pearson's Chi-squared test
data: spending_table
X-squared = 109, df = 5, p-value < 2.2e-16R
# 4: Standardized residuals
chi_spending$residuals |> round(2)
OUTPUT
High Low
Canada 3.24 -3.19
Colombia -3.14 3.08
Netherlands 2.60 -2.56
Other -2.82 2.77
United States 3.24 -3.19
Venezuela -3.14 3.08R
# Positive residuals in the "High" column mean that origin country has
# more high-spending rows than expected under independence.
Linear regression
In SPSS: Analyze > Regression > Linear. Move the dependent variable to “Dependent” and independent variables to “Independent(s)”.
Let’s predict average spending from stay nights and origin country:
R
reg_model <- lm(avg_spending_usd ~ avg_stay_nights + origin, data = visitors)
summary(reg_model)
OUTPUT
Call:
lm(formula = avg_spending_usd ~ avg_stay_nights + origin, data = visitors)
Residuals:
Min 1Q Median 3Q Max
-112.239 -7.704 2.216 12.417 58.083
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 234.210 33.037 7.089 1.24e-10 ***
avg_stay_nights 134.943 4.879 27.659 < 2e-16 ***
originColombia -184.753 12.670 -14.582 < 2e-16 ***
originNetherlands -619.305 17.829 -34.737 < 2e-16 ***
originOther -88.610 9.756 -9.082 4.05e-15 ***
originUnited States 114.139 6.600 17.294 < 2e-16 ***
originVenezuela -273.788 13.452 -20.354 < 2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 20.66 on 113 degrees of freedom
Multiple R-squared: 0.9937, Adjusted R-squared: 0.9933
F-statistic: 2962 on 6 and 113 DF, p-value: < 2.2e-16Reading regression output — SPSS comparison
The summary() output contains everything from the SPSS
regression output tables, but in a more compact format:
| SPSS table | R output section |
|---|---|
| Model Summary (R-sq) |
Multiple R-squared, Adjusted R-squared at
the bottom |
| ANOVA table (F-test) |
F-statistic at the very bottom |
| Coefficients table | The Coefficients: section |
| B (unstandardized) |
Estimate column |
| Std. Error |
Std. Error column |
| t |
t value column |
| Sig. |
Pr(>|t|) column |
Note: R does not give you standardized coefficients
(Beta) by default. To get those, scale your variables first with
scale(), or use the lm.beta package.
Reading R output with broom::tidy()
The raw R output is fine for interactive exploration, but it is hard
to export or combine with other results. The broom package
converts statistical output into tidy data frames — one row per term,
columns for estimate, standard error, test statistic, and p-value.
R
# Tidy the t-test result
tidy(t_result)
OUTPUT
# A tibble: 1 × 10
estimate estimate1 estimate2 statistic p.value parameter conf.low conf.high
<dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 -249 978. 1228. -16.2 1.21e-18 38.0 -280. -218.
# ℹ 2 more variables: method <chr>, alternative <chr>R
# Tidy the regression coefficients
tidy(reg_model)
OUTPUT
# A tibble: 7 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) 234. 33.0 7.09 1.24e-10
2 avg_stay_nights 135. 4.88 27.7 3.93e-52
3 originColombia -185. 12.7 -14.6 9.90e-28
4 originNetherlands -619. 17.8 -34.7 3.86e-62
5 originOther -88.6 9.76 -9.08 4.05e-15
6 originUnited States 114. 6.60 17.3 1.56e-33
7 originVenezuela -274. 13.5 -20.4 1.35e-39R
# Get model-level statistics (R-squared, F, etc.)
glance(reg_model)
OUTPUT
# A tibble: 1 × 12
r.squared adj.r.squared sigma statistic p.value df logLik AIC BIC
<dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 0.994 0.993 20.7 2962. 8.91e-122 6 -530. 1076. 1098.
# ℹ 3 more variables: deviance <dbl>, df.residual <int>, nobs <int>The tidy() output is a regular data frame, which means
you can filter it, arrange it, or export it to CSV — something that is
surprisingly difficult with SPSS output. This is extremely useful for
reporting: you can extract exactly the numbers you need and drop them
into a table or a plot.
This is a good time to reinforce the reproducibility advantage. In SPSS, if a reviewer asks you to re-run an analysis with a different subset, you have to click through the dialogs again. In R, you change one line of code and re-run the script.
Emphasize that broom::tidy() produces a data frame —
this means students can use all the dplyr verbs they learned in Episode
3 on their statistical results (filtering significant results, arranging
by p-value, etc.).
A complete analysis workflow
Let’s put everything together in a workflow that mirrors what you would do in SPSS, but entirely in code. Our research question: Does average spending differ by origin country, and what predicts higher spending?
R
# Step 1: Descriptive statistics by group
visitors |>
group_by(origin) |>
summarise(
n = n(),
mean_spending = mean(avg_spending_usd),
sd_spending = sd(avg_spending_usd),
mean_satisfaction = mean(satisfaction_score)
) |>
arrange(desc(mean_spending))
OUTPUT
# A tibble: 6 × 5
origin n mean_spending sd_spending mean_satisfaction
<chr> <int> <dbl> <dbl> <dbl>
1 United States 20 1228. 48.2 8.00
2 Canada 20 1139 63.0 7.90
3 Netherlands 20 978. 49.0 7.88
4 Other 20 850 64.2 7.48
5 Colombia 20 654 68.6 7.4
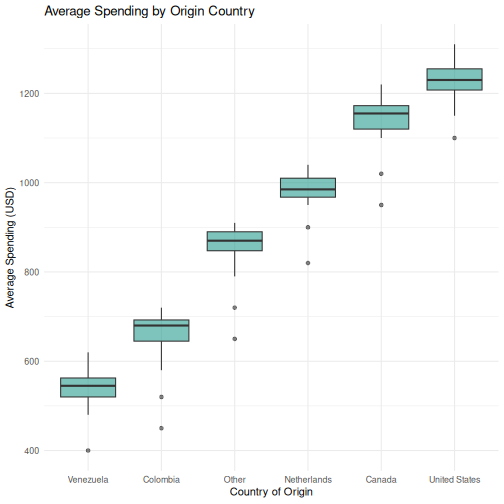
6 Venezuela 20 540 46.9 7.14R
# Step 2: Visualize the distribution
ggplot(visitors, aes(x = reorder(origin, avg_spending_usd, FUN = median),
y = avg_spending_usd)) +
geom_boxplot(fill = "#2a9d8f", alpha = 0.6) +
labs(title = "Average Spending by Origin Country",
x = "Country of Origin",
y = "Average Spending (USD)") +
theme_minimal()
R
# Step 3: ANOVA --- is there a significant difference?
spending_anova <- aov(avg_spending_usd ~ origin, data = visitors)
summary(spending_anova)
OUTPUT
Df Sum Sq Mean Sq F value Pr(>F)
origin 5 7262707 1452541 441.7 <2e-16 ***
Residuals 114 374890 3289
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1R
# Step 4: Post-hoc --- which groups differ?
tukey_results <- TukeyHSD(spending_anova)
tidy(tukey_results) |>
filter(adj.p.value < 0.05) |>
arrange(adj.p.value)
OUTPUT
# A tibble: 15 × 7
term contrast null.value estimate conf.low conf.high adj.p.value
<chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl>
1 origin Colombia-Canada 0 -485 -538. -432. 2.43e-14
2 origin Other-Canada 0 -289 -342. -236. 2.43e-14
3 origin Venezuela-Canada 0 -599 -652. -546. 2.43e-14
4 origin Netherlands-Colomb… 0 324. 272. 377. 2.43e-14
5 origin United States-Colo… 0 573. 521. 626. 2.43e-14
6 origin United States-Neth… 0 249. 196. 302. 2.43e-14
7 origin Venezuela-Netherla… 0 -438. -491. -386. 2.43e-14
8 origin United States-Other 0 377. 325. 430. 2.43e-14
9 origin Venezuela-Other 0 -310 -363. -257. 2.43e-14
10 origin Venezuela-United S… 0 -687. -740. -635. 2.43e-14
11 origin Other-Colombia 0 196 143. 249. 6.49e-14
12 origin Netherlands-Canada 0 -161. -213. -108. 2.75e-13
13 origin Other-Netherlands 0 -128. -181. -75.9 1.83e- 9
14 origin Venezuela-Colombia 0 -114. -167. -61.4 9.17e- 8
15 origin United States-Cana… 0 88.5 35.9 141. 5.04e- 5R
# Step 5: Regression --- what predicts spending?
spending_model <- lm(avg_spending_usd ~ avg_stay_nights + origin + year,
data = visitors)
tidy(spending_model) |>
mutate(across(where(is.numeric), ~ round(., 3)))
OUTPUT
# A tibble: 8 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) -7888. 2707. -2.91 0.004
2 avg_stay_nights 131. 4.92 26.6 0
3 originColombia -194. 12.6 -15.4 0
4 originNetherlands -605. 17.9 -33.9 0
5 originOther -94.8 9.65 -9.82 0
6 originUnited States 113. 6.38 17.8 0
7 originVenezuela -284. 13.4 -21.1 0
8 year 4.03 1.34 3 0.003R
# Step 6: Model fit
glance(spending_model) |>
select(r.squared, adj.r.squared, p.value, AIC)
OUTPUT
# A tibble: 1 × 4
r.squared adj.r.squared p.value AIC
<dbl> <dbl> <dbl> <dbl>
1 0.994 0.994 6.69e-122 1069.This entire analysis — from descriptives to regression — is captured in a script that you can re-run at any time. If your data updates or a reviewer requests changes, you modify the code and run it again. No clicking through dialog boxes.
Challenge 1: Complete analysis workflow
Conduct a full analysis to answer this question: Do satisfaction scores differ between high-season (Q1 and Q4) and low-season (Q2 and Q3) quarters?
Your workflow should include:
- Create a new variable
seasonthat classifies Q1/Q4 as “High” and Q2/Q3 as “Low” - Compute descriptive statistics (mean and SD of
satisfaction_score) for each season - Create a boxplot comparing satisfaction scores by season
- Run an independent-samples t-test
- Use
broom::tidy()to extract the results into a clean table
R
# Step 1: Create the season variable
visitors_season <- visitors |>
mutate(season = if_else(quarter %in% c("Q1", "Q4"), "High", "Low"))
# Step 2: Descriptive statistics
visitors_season |>
group_by(season) |>
summarise(
n = n(),
mean_satisfaction = mean(satisfaction_score),
sd_satisfaction = sd(satisfaction_score)
)
OUTPUT
# A tibble: 2 × 4
season n mean_satisfaction sd_satisfaction
<chr> <int> <dbl> <dbl>
1 High 60 7.71 0.382
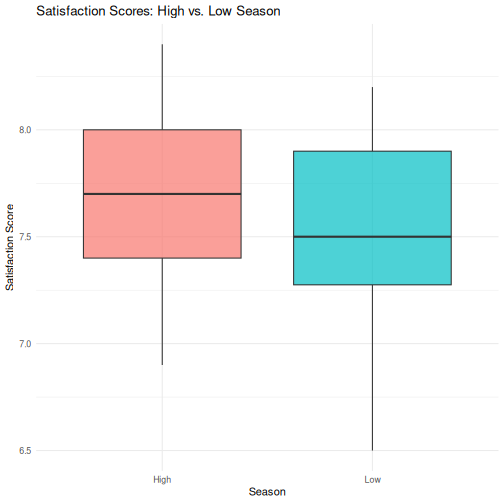
2 Low 60 7.55 0.405R
# Step 3: Boxplot
ggplot(visitors_season, aes(x = season, y = satisfaction_score,
fill = season)) +
geom_boxplot(alpha = 0.7) +
labs(
title = "Satisfaction Scores: High vs. Low Season",
x = "Season",
y = "Satisfaction Score"
) +
theme_minimal() +
theme(legend.position = "none")
R
# Step 4: T-test
season_t <- t.test(satisfaction_score ~ season, data = visitors_season)
season_t
OUTPUT
Welch Two Sample t-test
data: satisfaction_score by season
t = 2.3194, df = 117.6, p-value = 0.0221
alternative hypothesis: true difference in means between group High and group Low is not equal to 0
95 percent confidence interval:
0.02436478 0.30896855
sample estimates:
mean in group High mean in group Low
7.713333 7.546667 R
# Step 5: Tidy results
tidy(season_t)
OUTPUT
# A tibble: 1 × 10
estimate estimate1 estimate2 statistic p.value parameter conf.low conf.high
<dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 0.167 7.71 7.55 2.32 0.0221 118. 0.0244 0.309
# ℹ 2 more variables: method <chr>, alternative <chr>Challenge 2: Correlation and regression
Investigate the relationship between avg_stay_nights and
avg_spending_usd:
- Run
cor.test()to get the Pearson correlation and p-value - Create a scatterplot with a linear trend line
(
geom_smooth(method = "lm")) - Fit a linear regression predicting
avg_spending_usdfromavg_stay_nights - Use
tidy()andglance()to extract the results - Interpret: for each additional night of stay, how much does spending increase?
R
# Step 1: Correlation
cor.test(visitors$avg_stay_nights, visitors$avg_spending_usd)
OUTPUT
Pearson's product-moment correlation
data: visitors$avg_stay_nights and visitors$avg_spending_usd
t = 8.0872, df = 118, p-value = 6.065e-13
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
0.4680186 0.7013374
sample estimates:
cor
0.5971648 R
# Step 2: Scatterplot
ggplot(visitors, aes(x = avg_stay_nights, y = avg_spending_usd)) +
geom_point(aes(color = origin), alpha = 0.7, size = 2) +
geom_smooth(method = "lm", color = "black", linewidth = 0.8) +
labs(
title = "Relationship Between Length of Stay and Spending",
x = "Average Stay (Nights)",
y = "Average Spending (USD)",
color = "Origin"
) +
theme_minimal()
OUTPUT
`geom_smooth()` using formula = 'y ~ x'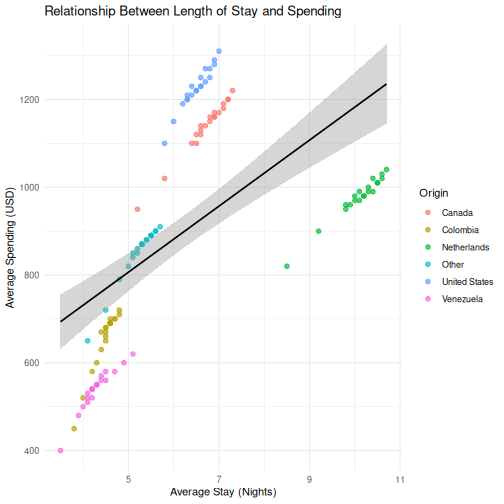
R
# Step 3: Regression
stay_model <- lm(avg_spending_usd ~ avg_stay_nights, data = visitors)
# Step 4: Extract results
tidy(stay_model)
OUTPUT
# A tibble: 2 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) 430. 60.8 7.07 1.16e-10
2 avg_stay_nights 75.3 9.31 8.09 6.06e-13R
glance(stay_model)
OUTPUT
# A tibble: 1 × 12
r.squared adj.r.squared sigma statistic p.value df logLik AIC BIC
<dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 0.357 0.351 204. 65.4 6.06e-13 1 -807. 1621. 1629.
# ℹ 3 more variables: deviance <dbl>, df.residual <int>, nobs <int>R
# Step 5: Interpretation
# The coefficient for avg_stay_nights tells us:
# for each additional night of stay, average spending increases by
# approximately the "estimate" value in USD, holding all else constant.
- Every SPSS statistical test has a direct R equivalent, usually in a single function call
- R output is more compact than SPSS —
broom::tidy()converts it to a clean table - The workflow in R is: load data, run test, extract results, visualize — all in a script
